| Breast Cancer Gene-Expression Miner v5.0
(bc-GenExMiner v5.0) | |  |
Tutorial - Gene correlation targeted analysis
|
|
Gene correlation targeted analysis permits to know the linear dependence between each pair of the 2 to 20 submitted genes.
|
|
|
| Step one |
Criterion 1:
gene expression data
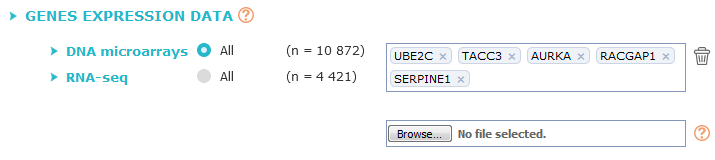
First choose the data source (All DNA microarrays or All RNA-seq), then fill the textbox with at least two and up to twenty actualised* gene symbols.
Gene symbols must be entered separately and validated one at a time (e.g.: SERPINE1 [enter]
UBE2C [enter] TACC3...).
A second way of requesting can be used by uploading your own gene list within a file.
Store your genes in a text file, with the text separator of your choice : tabulation, carriage return, comma or space. Note that only gene symbols will be taken into account.
Then upload your file with the second text box by clicking on "Browse" (here "Parcourir") button.
The screen capture have been made with a english browser, your own browser will use your personnal langage settings.
*: see actualised web databases (e.g.:
Ensembl,
GeneCards,
HGNC,
NCBI Gene...)
You can delete each gene symbol independently by clicking on the cross at the right side of the name, or delete whole list by clicking on trash bin (right side of the box).
|
|
Criterion 2:
criterion for studied population
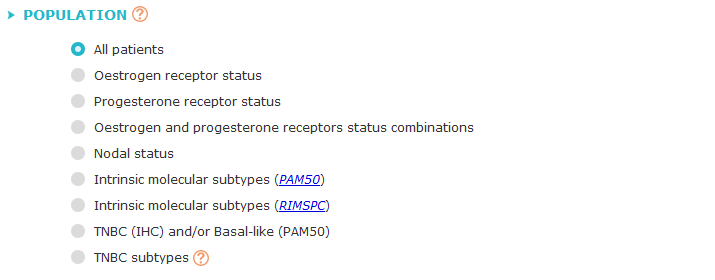
Choose one of the population characteristics (all patients, oestrogen receptor,
progesterone receptor, oestrogen and progesterone receptor combinations, nodal status,
intrinsic molecular subtypes [from PAM50], intrinsic molecular subtypes [from RIMSPC],
TNBC subtypes or basal-like/TNBC status) of the cohorts to be explored.
These datasets are retrieved from
annotated
transcriptomic data.
|
|
Last criterion: output figure
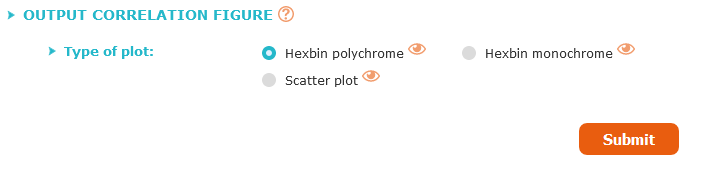
Correlation plots can be visualized and saved in several formats: hexbin plot, with a polychrome or monochrome representation, or "classic" scatterplot.
With the hexbin plot, the density of patients at the same point is represented by a different colorimetry, this colored hexagon adds additional information to the figure.
To optimize calculation time, only one type of figure will be generated, all are available but only one type at a time.
You have to choose which figure you will get at the results page, by clicking on the appropriate button.
You can preview the type of figure by clicking on the eye logo, a sample will be displayed.
Once all the criteria have been chosen, click on "Submit".
|
Step two |
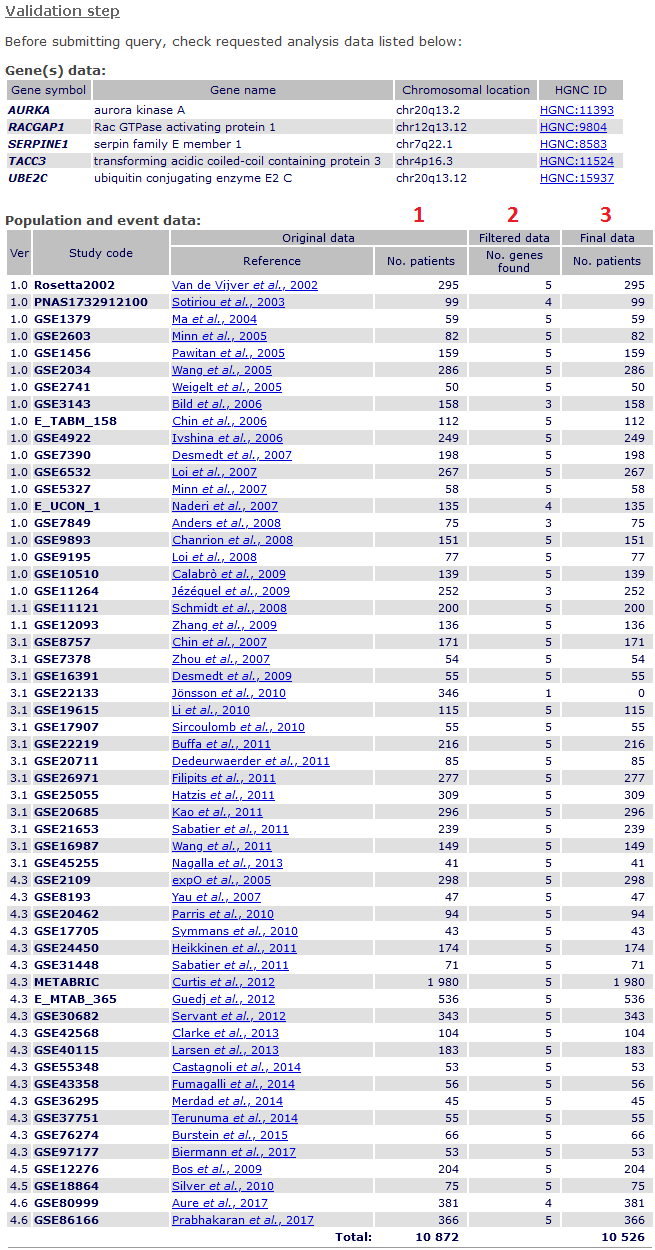
After submission, a validation page shows detailed information about:
- tested gene(s),
- patients from original studies tested,
1 complete data before filtering;
2 number of genes found;
3 patients finally analysed (if no missing genomic data).
|
|
|
After visualising the validation screen and reading the summary, at the bottom of the page, you can then choose to validate or cancel your submission according
to these intermediate descriptive data summarized at the bottom of the page.
- "Start analysis" will launch
statistical analyses
with the chosen gene and direct you to targeted correlation analysis result page.
- "Cancel" will redirect you back to previous screen, and offer you to choose a new gene or modify your list of genes.
|
|
|
|
The results page offers two ways of displaying the figures.
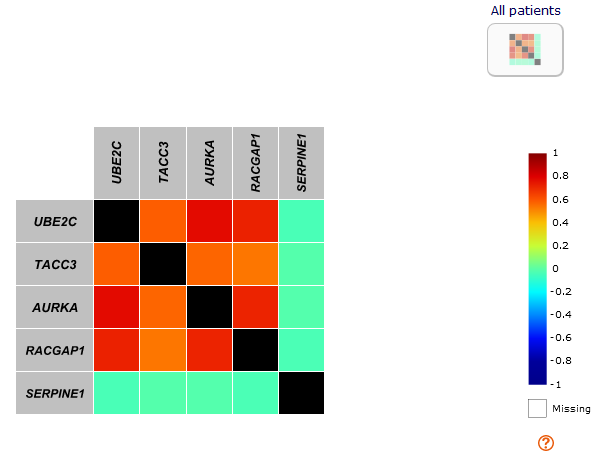
First, the "old method" is kept with a solid square
correlation map
corresponding to the analysis with all patients.
The two sides of the diagonal (in black) are symmetrical and contain only the color of the correlation coefficient value (according to the scale value legend).
Help to interpret correlation coefficient value is available by clicking on the "?" symbol below the colour scale.
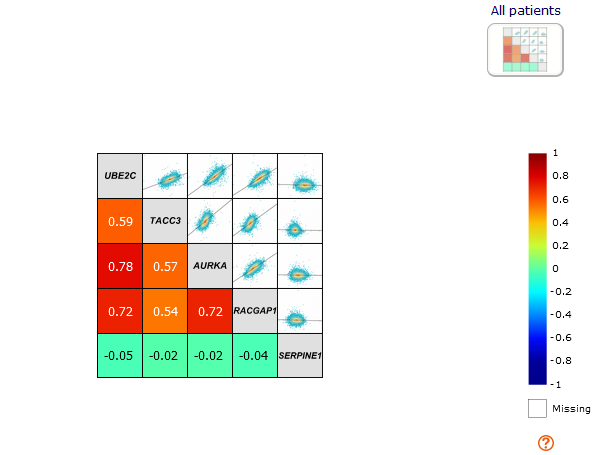
Then the "new method" is proposed with a
correlation map
with two half squares corresponding to the analysis with all patients.
The diagonal contains the gene symbol, so the names are inside the map.
The lower left part of the diagonal contains the colored tiles of the correlation coefficient value (according to the scale legend)
and the value itself.
Hovering over the tiles will bring up a pop-up window that adds more features to the reading of your results.
Next, the top right side of the map displays the correlation graph for each pair of genes, based on the output figure chosen at the beginning of the analysis process.
Again, hovering over the cells will add more information, and the tile is clickable and offers a full screen of the gene pair correlation results.
Correlation maps corresponding to analysis among patients with respectively:
- all patients,
- positive or negative oestrogen receptor (ER) status,
- positive or negative progesterone receptor (PR) status,
- combination of ER/PR statuses,
- positive or negative nodal (N) status,
- patients of the five molecular subtypes as determined by the PAM50,
- patients of the four molecular subtypes as determined by the RIMSPC,
- patients determined by basal-like PAM50 subtyped and/or triple-negative breast cancer typed by immunohistochemistry,
- and finally triple-negative breast cancer subtypes patients (C1, C2 and C3 subtypes)
can be displayed by clicking on the corresponding button below.
Depending on population criterion chosen before, respectively like the first request page, the "above buttons bar" will be presented like following capture :
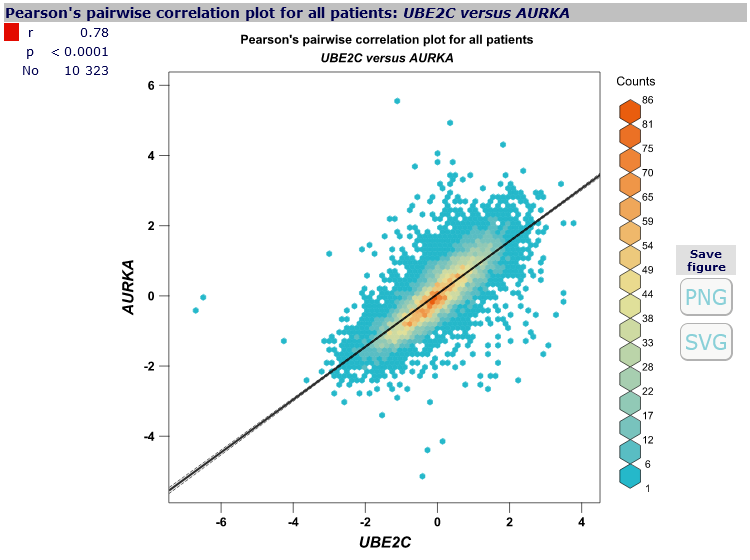
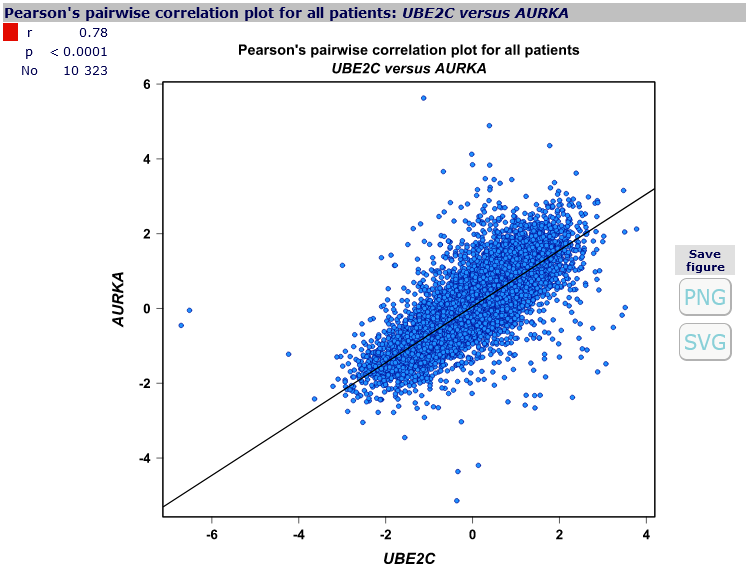
For each pair of genes,
correlation plot
can be displayed, along with correlation coefficient value,
corresponding p-value and number of patients involved, by clicking on the corresponding cell in the correlation map.
Depending on the choice made in the first step, different figures will be displayed:
- hexbin polychrome,
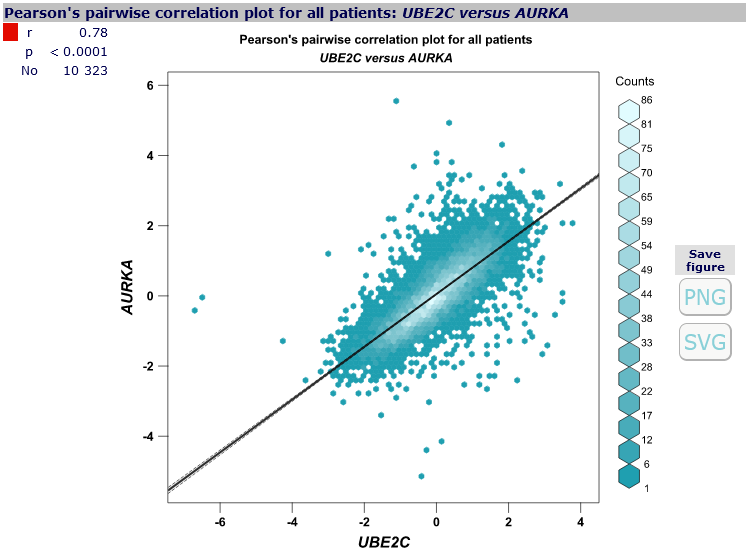
- hexbin monochrome,
- scatter plot,

|
Correlation maps can be saved in "PNG" ([portable network graphics] an universal and easy to use format) or
"SVG" ([scalable vector graphics] a lossless image format figure that allows edit viewing and printing settings, as you wish, for your research article) format. |

|
Tables with all detailed values (correlation coefficient, p-value, number of patients) can be saved in csv [comma separated values] format.
|
|
|
|